Pannexin 3 has distinct structural features in comparison to other isoforms
The PANX3 isoform shares 42% sequence identity with PANX1 (Supplementary Fig. 1) and 27% identity with PANX2. To understand the architectural differences among PANX isoforms, we purified the full-length human PANX1 and 3 isoforms and elucidated the cryo-EM structures to an overall resolution of 3.75âà and 3.9âà , respectively (Supplementary Figs. 2 and 3) The PANX3 structure was determined in the presence of ATP (1âmM) and high K+ (100âmM; Fig. 1a, b, Supplementary Fig. 3, and Table 1). Similar to PANX1, high extracellular potassium is also speculated to open PANX3 channels30. We used 100âmMâK+ during final purification of both PANX1 and PANX3 in an attempt to capture an open conformation of the channels. While we successfully determined the structure of PANX1WT, for structural comparison, we utilized previously published PANX1 (PDB ID-6WBF) structure coordinates due to its higher resolution26. The density at the pore and transmembrane helices in PANX3 was sufficient to model most of the side chains in these regions (Supplementary Fig. 4). The intracellular helices (160-185) and the C-terminus (373-392) lack clear densities, likely due to inherent flexibility in this region. Density for bound ATP molecules that would allow unambiguous assignment, at this resolution, was not observed. The PANX3 retains the heptameric oligomer assembly that can be partitioned into an extracellular domain (ECD), transmembrane domain (TMD), and intracellular domain (ICD), similar to PANX1WT and PANX2 (Fig. 1a, b). Unlike other large-pore ion-channels like CALHMs31, the PANX isoforms do not display heterogenous oligomeric associations32. The PANX3 channel is 8âà wider than the PANX1 at the cytosolic face with similar transmembrane length in the ordered regions of the channel (Fig. 1c). The channel width of PANX3 at the extra and intracellular faces is similar to PANX2 (Fig. 1c).
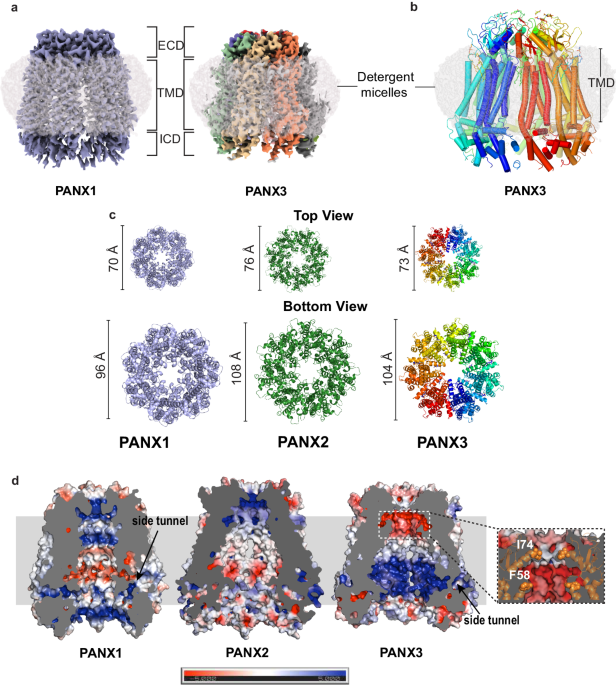
a The cryo-EM map for PANX1 (blue) and PANX3 (protomers colored individually) viewed parallel to the membrane plane and surrounded by the detergent micelle (gray). b The modeled structure of PANX3 displaying heptameric organization and embedded in a detergent micelle that marks the transmembrane boundary. c Top (extracellular) and bottom (cytosolic) views of PANX1, 2 and PANX3 exhibiting differences in the dimensions in PANX isoforms at the extracellular and intracellular faces of the channel. d Sagittal section of surface electrostatics of PANX1 (PDB ID: 6WBF), PANX2 (PDB ID: 7XLB), and PANX3 colored according to potential from â5 (red) to +5 (blue) (kBTecâ1) viewed parallel to the membrane plane. The inset shows the position of the residues I74 and F58 (orange spheres) forming first and second constrictions, respectively in PANX3.
The topology of PANX3 protomers is similar to PANX1WT with differences in the TM1 and the extracellular loops. The topology comprises four transmembrane helices and two extracellular loops (EL). Two disulfide bonds (SS1 and SS2) stabilize the extracellular domain, C66-C261(SS1) and C84-C242 (SS2), that form between EL1 and EL2 (Supplementary Fig. 5a, b). The superposition of two protomers (PANX1 vs. PANX3) yields a root mean square deviation (rmsd) of 3.2âà for the 302âCα atoms aligned. There is also a significant alteration of surface charge within PANX3 compared to PANX1WT and PANX2 near the surface lining the solvent-accessible channel vestibule (Fig. 1d).
N-linked glycosylation at the N255 position in the EL2 of PANX1 was implicated in preventing the formation of gap junctions26,33. A substitution at this site (N255A) led to the formation of a mixture of gap junctions and hemichannels26. In the PANX3 structure, we observed density for N-acetylglucosamine (NAG) at the predicted glycosylation site (N71) in the first extracellular loop that is much closer to the pore, in comparison to PANX1 (Supplementary Fig. 5c, d). The N-glycosylation in PANX2 is also localized to the EL1 region at N86 position that coincides with PANX3 N-glycosylation site (Supplementary Fig. 1) although the structure of PANX2 in a recent report does not reveal the site due to a likely disorder in the vicinity17.
In PANX3, we do not observe density for residues 1-24 in N-terminus. It is therefore unclear if the N-terminus lines the pore and plays a role in maintaining the rigidity of the transmembrane domain of the heptamer similar to PANX1. Towards the C-terminus, we did not observe the density for the last twenty residues (373â392). The C-terminus is comparatively shorter than the corresponding stretch of PANX1 and lacks a caspase cleavage site to facilitate ATP release, as observed in PANX1. Moreover, ATP release has been observed in PANX3 in presence of high extracellular potassium even without the deletion of C-terminus of PANX330. To test the functional effects of the C-terminus in affecting channel properties, we designed a deletion construct (PANX3C-del) of PANX3 lacking the last 22 residues (372â392). We performed electrophysiology experiments and found no discernible change in current associated with the deletion of the C-terminus when compared with PANX3WT, although the surface expression of the deletion mutant was higher than PANX3WT (Supplementary Fig. 6a, b). This observation led us to suggest that the short C-terminus of PANX3 may not play a significant role in channel opening and PANX3 may have alternate mechanisms for channel-gating.
PANX3 has a wider pore among pannexins, and pore mutants alter channel properties
The residue lining the constriction point in PANX1, W74, is replaced by isoleucine or valine in different orthologues of PANX3 (Fig. 2a, b and Supplementary Fig. 6c). The human PANX3âs pore is lined by two residues, I74 and R75, with a width of 13.2âà at the narrowest point created by I74 (Fig. 2b). The primary constriction created by the W74 residue in PANX1WT results in a pore diameter of 12âà . The presence of I74 in PANX3 instead of W74 widens the pore to a diameter of 13.2âà . The cation-Ï interaction between W74 and R75 in PANX1WT is eliminated in PANX3 due to the replacement of W74 with I74. R75 of one PANX3 protomer interacts with D81 of the other protomer through a salt bridge (Fig. 2d, f, h). Residues in the same position in the pore in PANX2 are substituted with R89 and D90 that are drastically different from residues observed in PANX1 and PANX3. R89 in PANX2 is suggested to form highly cationic pore at the channel entrance (Fig. 2e). However, unlike PANX1 and PANX3, the main pore residues, R89 and D90 in PANX2, do not engage in interprotomeric interactions (Fig. 2e, g)17.
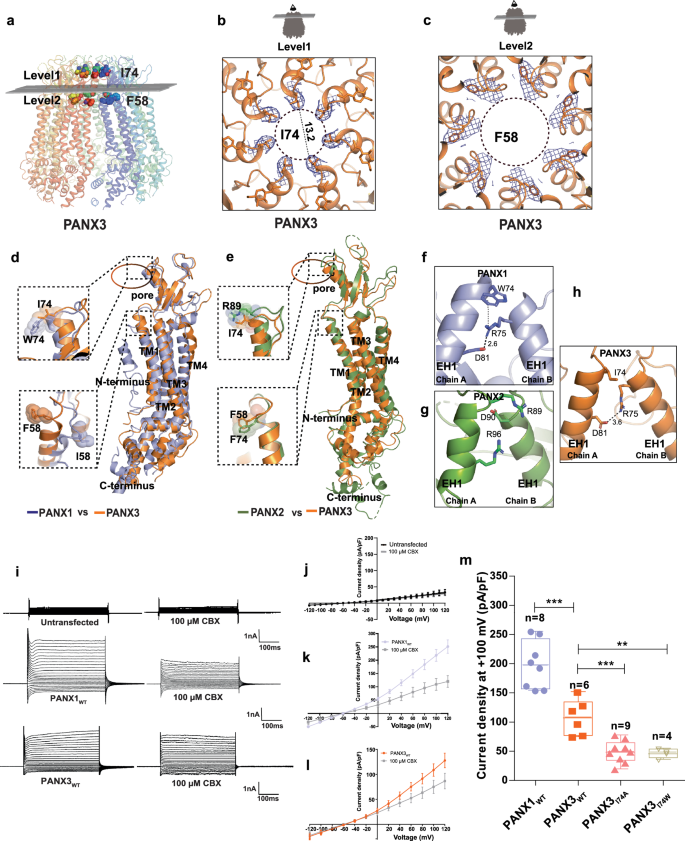
a Transverse sections of PANX3 at two distinct levels are presented, illustrating the locations of two constrictions. Specifically, I74 and F58 in PANX3 contribute to the formation of the first and second constrictions, respectively. b A close-up of the first constriction seen from above in PANX3 formed by I74. The density for the residue (I74) at 7.5 Ï is shown. c A close up of the second constriction formed by F58 in PANX3. The density for the residue (F58) at 7.5 Ï is shown. d Superposition of PANX1 and PANX3 displaying the position of residues 58 and 74 with a rmsd of 3.2âà for 302âCα atoms, e Superposition of PANX2 (PDB ID: 7XLB) and PANX3 exhibits the differences in the pore residues, R89 and I74 in PANX2 and PANX3 respectively with a rmsd of 5.4âà for 288 atoms aligned; F74 in PANX2 acquires similar position as F58 in PANX3. f Cation-pi interaction(dotted lines) between W74 and R75 in PANX1 (PDB ID: 6WBF) is lost in PANX3. g Hydrogen bond interaction between R75 and D81 in PANX1 is also observed in PANX3 similar to PANX1. All the distances depicted in the figure are in angstroms (à ). h Pore residue (R89, D90) in PANX2 do not form any interactions with neighboring residues unlike PANX1 and PANX3 iâl, Representative traces for whole-cell current for HEK293 untransfected cells(mock) and HEK293 cells expressing PANX1WT and PANX3 with and without CBX (100âµM) application. Current density-voltage plot for untransfected, PANX1 and PANX3 in presence and absence of CBX. Each point represents the mean of nâ=â5 (untransfected), nâ=â4 (PANX1) and nâ=â6 (PANX3) individual recordings, and the error bar represents SEM. m Current density is plotted for the PANX1WT (nâ=â8) and PANX3WT (nâ=â6) and its mutants, PANX3I74A (nâ=â9), PANX3I74W (nâ=â4), the error bar represents SEM. n represents the number of cells used for independent recordings; a two-tailed unpaired t-test is used for calculating the significance, ***pâ<â0.001; n.s., not significant, PANX1WT vs PANX3WT (P valueâ<â0.0001), PANX3WT vs PANXI74A (P valueâ=â0.0005), PANX3WT vs PANXI74W (P valueâ=â0.0047), The whiskers represent minimum and maximum value, the left edge of the box represents 25% quartile and the right edge represents 75% quartile, the middle line represents median. Box plot statistics for PANX1WT are, minimum (154.5), 25% percentile (156.9), median (198.0), 75% percentile (242.8), maximum (255.6), for PANX3WT, minimum (75.03), 25% percentile (77.00), median (108.6), 75% percentile (134.4), maximum (152.0), for PANX3I74A, minimum (49.96), 25% percentile (56.77), median (64.21), 75% percentile (77.97), maximum (97.97), for PANX3I74W, minimum (36.24), 25% percentile (38.92), median (48.44), 75% percentile (53.84), maximum (55.14), raw traces and the IV curve for the PANX3 mutants are presented in Supplementary Fig. 12.
Moreover, S70 and Q76 between two protomers of PANX3 form a hydrogen bond resulting in interprotomeric interactions leading to a stable heptamer. Apart from these interactions, there seem to be minimal intersubunit interactions across the ECD and TMD regions between the protomers (Supplementary Fig. 6e). The gap between the TMs 2 and 4 of adjacent protomers just beneath the ECD is occupied by lipid-like density into which we have modeled a phospholipid corresponding to the upper leaflet of the bilayer, 1-palmitoyl-2-oleoylphosphatidylethanolamine (POPE), for each protomer (Supplementary Fig. 5e). The phospholipid could be enhancing the interactions between the protomers of PANX3 akin to the phospholipid interactions observed in the CALHM1 channels34. However, besides this lipid density we did not observe any other prominent density for lipid at the inner leaflet of the bilayer likely due to the lower resolution of this structure compared to other channels among large-pore ion channels. As a consequence, we could observe a gap between subunits that is large enough to allow the passage of ions. The presence of a side-tunnel suggested by Ruan et al. in PANX1WT is prominent in PANX3 with a 6.9âà separation between TM2 and CTH1 compared to 6.3âà in PANX1WT, suggesting that the lateral portal hypothesis may also hold true for PANX3 (Supplementary Fig. 6f). Although we did not observe a lipid density around this region that could occlude this site, a higher resolution structure of PANX3 might reveal greater details about the solvent accessibility around this portal. This region is occluded with a distance of 2.6âà between TM2 and CTH1 in the case of PANX2 and is unlikely to support the entrance of ions from this portal in PANX2 in the conformation, reported from the model (Supplementary Fig. 6f).
The structural comparison of PANX3 with PANX1WT reveals an additional constriction in PANX3 below the primary channel pore. The residues at the end of TM1, 58-60 comprising residues F58, S59, and S60 form a prominent second constriction at the neck region in PANX3 compared to PANX1WT (Fig. 2a, c). The linker between TM1 and TM2 adopts a clear α-helical conformation in PANX3 and PANX2, instead of a loop observed in PANX1WT, constricting the vestibule in the region, thus allowing PANX3 to have an additional vestibule beneath the pore (Fig. 2a, d). The residues F58-S59-S60 line the second constriction point facing the pore in PANX3 and are part of the PANX3 sequence that is variable between PANX1, 2 and 3 (Supplementary Fig. 6d). In contrast, I58 residue in PANX1WT participates in hydrophobic interactions between TMs 1 and 2. Despite sequence variation between PANX2 and PANX3, the F74 in PANX2 forms a similar motif that can allow the demarcation of the vestibule into two regions even in PANX2 isoform (Fig. 2e). The diameter at this constriction point is 21âà in PANX3 compared to 30âà in PANX1WT and demarcates the boundary between the anionic surface of the upper compartment compared to the amphiphilic lower compartment in PANX3 (Fig. 1d). Additionally, an annulus of seven uncharacterized densities is observed at this second constriction point in PANX3 that might further contribute to restricting access at this region (Supplementary Fig. 6g).
Given the role of PANX 1 and 3 isoforms in ATP release activity9,30, we analyzed ATP interactions with the channel through binding analysis of ATP-γS with PANX isoforms utilizing microscale thermophoresis (MST). Both ATP and ATP-γS displayed similar affinities when tested using PANX1WT that encouraged us to use the non-hydrolysable analog of ATP for MST binding assays (Supplementary Fig. 7a, b) The affinity of ATP analog ATP-γS for PANX3 was determined to be 69âµM compared to the 18âµM affinity observed for PANX1WT (Supplementary Fig. 7c). The ATP-γS binding studies suggest a weaker affinity of PANX3 for ATP-γS compared to PANX1. Although we observe a lower affinity of ATP-γS for PANX3, it is difficult to speculate the factors affecting the affinity, as a clear ATP binding site is yet to be detected in large-pore ATP channels such as CALHM, connexins and pannexins.
Furthermore, different pore-lining residues observed in PANX1 and PANX3 may dictate isoform-specific differences in channel properties as observed in connexins35. Despite earlier unsuccessful attempts to record currents from PANX324,25, we could observe currents in PANX3 heterologously expressed in HEK293 cells (Fig. 2i). Whole-cell current elicited by PANX3 shows similar I-V profile as PANX1 with outwardly rectifying currents at positive potential (Fig. 2i-l). We observe weaker CBX sensitivity in PANX3, in comparison to PANX1 (Fig. 2k, l, Supplementary Fig. 7d). The ECD region, particularly loop1, surrounding the pore was previously implicated as being important for CBX interactions25. The substitutions among pore lining residues observed in PANX3 could be a reason for the reduced sensitivity to CBX (Supplementary Fig. 7d). This also reinforces the observation of PANX1 pore substitutions, W74I and W74A, that lack sensitivity to CBX inhibition25. The ability of CBX to interact with PANX3, albeit weakly, despite a wider pore, strengthens the idea that CBX interactions at extracellular domain can modulate channel activity instead of directly acting as an asymmetric pore blocker. Alternate uncharacterized sites for CBX interactions can exist within PANX isoforms given its sterol-like chemical structure that modulates channel activity. This is reinforced by the lack of saturation in CBX interactions when tested using binding experiments (Supplementary Fig. 7e, f). The structural and the vestibule surface electrostatic differences in the pores observed between the three isoforms PANX1, 2 and 3 can therefore influence their respective channel activities and, consequently, their physiological effects (Fig. 1d). A notable difference between PANX1 and PANX3 is the pore-lining residue, I74 in PANX3 as opposed to W74 in PANX1. To unravel the consequences of this difference, we substituted I74 to a tryptophan (I74W) and alanine (I74A), respectively and observed the impact of these alterations on channel function, in PANX3. We observed that substitutions at this site are not tolerated and lead to a loss of any measurable current and a reduced surface expression among PANX3 substitutions (Fig. 2m and Supplementary Fig. 6b). This is unlike PANX1 where W74I substitution retained PANX1WT like currents25.
PANX1 pore substitutions mimic pore residues of PANX2
As observed in the PANX3I74W mutant, a single mutation at the pore led to an inactive channel. Moreover, single substitutions of W74R and R75D in the pore region yielded incorrectly assembled PANX126. In order to observe if a dual substitution of both the residues (W74, R75) by charged amino acids could compensate for this behavior, we created a PANX1 double mutant (PANX1WR/RD) by substituting W74 and R75 with arginine and aspartate, respectively (Fig. 3a). Incidentally, these substitutions are similar to the pore residues of PANX2 revealed by multiple sequence alignment and structural comparison with PANX2 (Supplementary Fig. 1 and Fig. 3b, c). Surprisingly, we obtained a minor fraction of well-assembled PANX1WR/RD particles that allowed reconstruction of the PANX1WR/RD structure to a resolution of 3.2âà (Table 1 and Supplementary Fig. 2b). Given the high resolution of this reconstruction, the densities along the pore and vestibule were clear for modeling residue sidechains (Fig. 3c). Whilst the global structure resembles the PANX1WT, the pore diameter was reduced to 9.3âà compared to 12.1âà in PANX1WT (Fig. 3b). The presence of R74 side chain facing the pore renders the pore of PANX1WR/RD highly cationic as compared to PANX1WT and the pore diameter resembles PANX2 (Fig. 3dâf)17. The main interactions at the pore in PANX1WT are W74-R75 cation-pi interaction within the subunit and R75-D81 interaction between two subunits (Fig. 2f). As these residues were mutated, the interactions are lost in the PANX1WR/RD (Fig. 3c). Interestingly, these interactions are vital to stabilize the heptamer, and the charge reversal mutant R75E does not form stable heptamers26. Instead, in PANX1WR/RD, D75 of one protomer interacts with R74 of another protomer to stabilize the structure (Fig. 3a, c). Apart from the alterations in the pore size, there are no substantial differences induced by the pore mutation, and the overall structure still bears resemblance to PANX1.
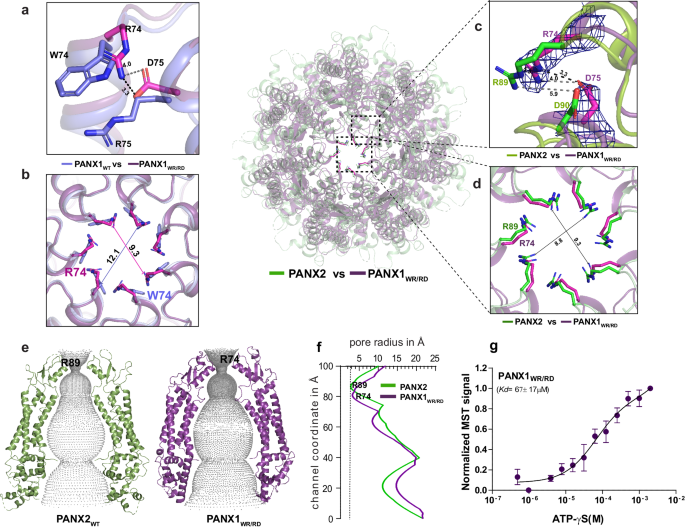
a The superposition of PANX1 (blue) and PANX1WR/RD (violet) illustrates that in PANX1WR/RD, R74 and D75 serve as pore residues, contrasting with W74 and R75 in PANX1WT. b A cross-section of superposed PANX1WT and PANX1WR/RD displays the reduction of pore. c The superposition of PANX1WR/RD and PANX2 unveils the pore residues in PANX2, with the density for R74 and D75 in PANX1WR/RD contoured at 9.0 Ï. d A cross-section of superposed PANX2 and PANX1WR/RD displays the position of arginine at the pore. The distances depicted in the figure are in angstroms (à ). e, f The hole profile displays similar pore radius for PANX1WR/RD and PANX2, The line represents the minimum radius for PANX2 at 2.7âà formed by the first constriction (R89) in PANX2 channel. Units of both X and Y-axes are in angstroms (à ). g The binding affinity for the PANX1WR/RD was determined to be 67â±â17âµM, nâ=â3 independent experiments, error bar represents S.D.
We analyzed the differences of ATP-γS interactions with PANX1WR/RD and PANX1WT via MST binding. This revealed a reduced Kd value of 67âμM in PANX1WR/RD compared to ATP-γS binding affinity for PANX1WT (18âµM), suggesting that ATP interactions are moderately affected by these pore substitutions (Fig. 3g and Supplementary Fig. 7b).
We also investigated the physiological consequences of these modifications on the channel behavior. The data revealed a two-fold reduction in current density in PANX1WR/RD compared to PANX1WT but retained an active channel despite the constriction of the pore radius (Fig. 3b, dâf and Supplementary Fig. 8a). In addition, we observed a comparatively higher surface expression for the mutant than PANX1WT (Supplementary Fig. 8b). We also observed a decreased inhibition of PANX1WR/RD currents with CBX compared to PANX1WT since the primary binding site for CBX, W74, was substituted to an arginine (Supplementary Fig. 7d). It is rather interesting that CBX retains minimal interactions with the PANX1WR/RD despite major substitutions in the pore. This further indicates the presence of alternate sites for CBX binding that could modulate channel gating. Additionally, the conductance density shows a fall in channel conductivity for PANX1WR/RD, although normalized G-V curve plotted for the double mutant displays an unaltered voltage sensitivity (Supplementary Fig. 8c and Fig. 4eâg). From this, we can infer that the dual pore substitutions done in PANX1 do not influence the voltage-dependent conductance property of the channel.
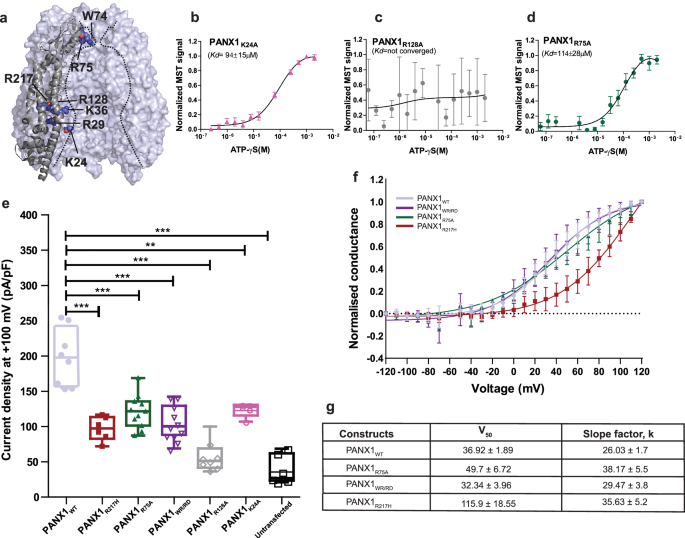
a The position of the mutants selected for ATP-γS binding and whole patch clamp experiments is displayed. b Microscale thermophoresis profile for PANX1R24A shows the binding of 94â±â15âµM. c MST data reveals a complete loss of binding in PANX1R128A mutant. d The binding affinity for the putative ATP binding site (R75) in PANX1 was determined to be 114â±â28âµM, suggesting that R75 is not the sole residue responsible for ATP-γS binding in PANX1, nâ=â3 independent experiments, error bar represents S.D. e Current density is plotted for the PANX1WT(nâ=â8) and the mutants, PANX1R75A(nâ=â11), PANX1R217H(nâ=â6), PANX1WR/RD(nâ=â11), PANX1K24A(nâ=â5) PANX1R128A(nâ=â7), and untransfected(nâ=â7) the error bar represents SEM. n represents the number of cells used for independent recordings; a two-tailed unpaired t-test is used for calculating the significance, ***pâ<â0.001; n.s., not significant, PANX1WT vs PANX1R75A (P valueâ<â0.0001), PANX1WT vs PANX1WR/RD (P valueâ<â0.0001), PANX1WT vs PANXR217H (P valueâ<â0.0001), PANX1WT vs Untransfected (P valueâ<â0.0001), PANX1WT vs PANXK24A (P valueâ=â0.0022), PANX1WT vs PANXR128A (P valueâ<â0.0001). The whiskers represent minimum and maximum value, the left edge of the box represent 25% quartile and the right edge represents 75% quartile, the middle line represents median. Box plot statistics are follows, for PANX1WT, minimum (154.5), 25% percentile (156.9), median (198.0), 75% percentile (242.8), maximum (255.6), for PANX1R217H are as follows, minimum (71.99), 25% percentile (82.77.), median (97.37), 75% percentile (113.5), maximum (116.9), for PANX1R75A, minimum (87.15), 25% percentile (101.4), median (121.7), 75% percentile (135.7), maximum (168.8), for PANX1WR/RD are as follows, minimum (69.0), 25% percentile (88.1), median (100.2), 75% percentile (129.4), maximum (142.5), for PANX1R128A, minimum (36.5), 25% percentile (40.1), median (49.3), 75% percentile (54.9), maximum (73.7), for PANX1K24A, minimum (107.0), 25% percentile (115.6), median (127.8), 75% percentile (130.7), maximum (130.9), for untransfected, minimum (20.9), 25% percentile (23.5), median (27.0), 75% percentile (62.0), and maximum (68.6), raw traces and the IV curve for the PANX1 mutants are presented in Supplementary Fig. 11. f Normalized conductance-voltage (GV-curve) plot for PANX1WT(nâ=â8), PANX1R75A(nâ=â6), and PANX1WR/RD(nâ=â4) suggests that the pore residues are not involved in the voltage sensitivity of the channel, conductance-voltage(GV-curve) plot for PANX1R217H exhibits a reduction in the voltage sensitivity of the channel, nâ=â4; the error bar represents SEM. g The normalized G-V values were fitted with the Boltzmann equation, and the voltage at which the half-maximal activation, V50, occurred along with slope factor, k, was calculated for all the constructs.
Our structural findings suggest that the pore radii are inherently linked to the specific residues lining this constriction. This correlation emphasizes the critical role of these residues in determining the dimensions and charge of the pore and, consequently, the functional properties of the PANX channels. It is evident from the functional studies of the three PANX isoforms, substitution of residues in the pore to resemble other isoforms helps alter the channel properties and the ability of inhibitors like CBX to interact with the modified pannexins18,25,26.
Vestibular cationic residues in PANX1 alter ATP interactions and channel properties
In addition to the pore substitutions among PANX isoforms, their ability to release ATP hinges on their capacity to facilitate ATP permeation through the channel prior to being released through the pore and it is likely to interact at different sites as it permeates through the channel. Given the distinct electrostatic surface properties within the vestibule of pannexins, we explored the effects of substituting positively charged residues along the vestibule in PANX1, like K24, R29 (N-terminus), K36 (TM1), R75 (EH1, pore), and R128 (TM2), that face the vestibule of the PANX1 channel (Fig. 4a). With the exception of R128, these vestibular residues are conserved in both PANX1 and PANX3 isoforms (Supplementary Fig. 1). Through alanine mutations, we systematically investigated the effects of these substitutions in PANX1 to decipher their impact on the channel function. The substitution of R29A and K36A resulted in excessive cell death and consequently, the purification of these PANX1 mutants was not feasible. The ATP-γS interaction with K24A substitution has a five-fold loss of affinity (Kd of 94âµM) compared to PANX1WT (Fig. 4b and Supplementary Fig. 7b). On the other hand, R128A (TM2) substitution reveals a complete loss of ATP-γS binding, indicating the importance of this region for PANX1 function (Fig. 4c). Moreover, R75 was suggested as a putative ATP binding site through alanine scanning mutagenesis36. We, therefore, mutated R75 to alanine and checked the binding affinity of PANX1R75A with ATP-γS. The MST data suggests a decrease in the binding affinity with PANX1WT but does not abolish it completely (Fig. 4d). Although we observe that R75 contributes to ATP-γS binding, it is not the sole residue for ATP binding as substitution of other positively charged residues also led to decreased ATP-γS binding (Fig. 4bâd). In all likelihood, multiple residues are involved in ATP interactions, and mutating these residues individually reduces ATP binding but does not abolish it entirely.
To further investigate the effects of these mutations, we performed whole-cell patch clamp studies. The PANX1R128A behaved similar to the untransfected cells and the loss of ATP-γS binding in PANX1R128A could be a result of improper oligomerization, as observed through the widened size-exclusion chromatography profiles (Supplementary Fig. 9). The N-terminus mutant K24A, exhibited weaker current density compared to the wild type in electrophysiology studies, which could be linked to its lower surface expression compared to PANX1WT (Fig. 4e and Supplementary Fig. 8b).
Moreover, electrophysiology data shows a two-fold decrease in the current density in PANX1R75A compared to PANX1WT, although surface expression for the mutant is significantly higher than the PANX1WT. Since the residue is involved in stabilizing interactions with W74 and interprotomeric interactions, R75A substitution could locally alter the interactions around the pore (Fig. 2f). Further, normalized G-V curve for R75A is comparable to wild type and reveals that the substitution of a pore residue, R75, although alters the channel properties but does not affect the voltage sensitivity of the channel (Fig. 4f, g) consistent with the findings with ZfPANX137. Similar consequences to arginine substitutions in the pore are observed with LRRC8A, where the channel maintains conductance with arginine substitution but loses anion selectivity38.
It is evident from these observations that the charge substitutions in the vestibule and the pore can influence ATP interactions with the PANX1 channel (Fig. 4bâd and Supplementary Fig. 8aâc).
Pannexin 1 germline mutant R217H leads to pore constriction
Encouraged by structural and functional alterations observed in both pore and vestibular mutants of PANX1, we explored the impact of a transmembrane germline mutation on the inherent properties of the channel. We elucidated the structure of PANX1R217H (TM3) substitution to characterize the effects of this germline mutant and to understand the basis of its defective channel properties. The arginine residue at the 217 position (TM3) is highly conserved among vertebrate PANX1 orthologues and is buried amongst TM helices and does not display solvent access unlike the substitutions studied earlier (Fig. 4a and Supplementary Fig. 10a). Consistent with the earlier reports15, we observed comparable expression of the PANX1R217H channel to PANX1WT (Supplementary Fig. 10b). The structure was determined by cryo-EM to a resolution of 3.9âà (Supplementary Fig. 2c and Table 1). We further improved the resolution of ECD and TMD domains to 3.77âà through focussed refinement (Supplementary Fig. 10c, d). However, we were unsuccessful in improving the resolution of ICD domain through this step. The superposition of PANX1WT and PANX1R217H mutant yielded a rmsd of 1.6âà for 288âCα atoms.
Despite the lower resolution of the structure, it resembles PANX1WT globally, with subtle changes in extracellular domain (ECD), extracellular helix 1 (EH1), and the intracellular domain (ICD). Extracellular loops and EH1 have an average shift of 1.4â1.6âà away from the pore. The W74 residue of PANX1R217H fits better into the density of its sidechain with a rotameric shift that coincidentally constricts the pore radius (Fig. 5a and Supplementary Fig. 10c). Despite a shift in the side chain position of R75, this residue retains interactions with the D81 of the adjacent subunit through a salt bridge, consistent with PANX1WT (Fig. 5b).
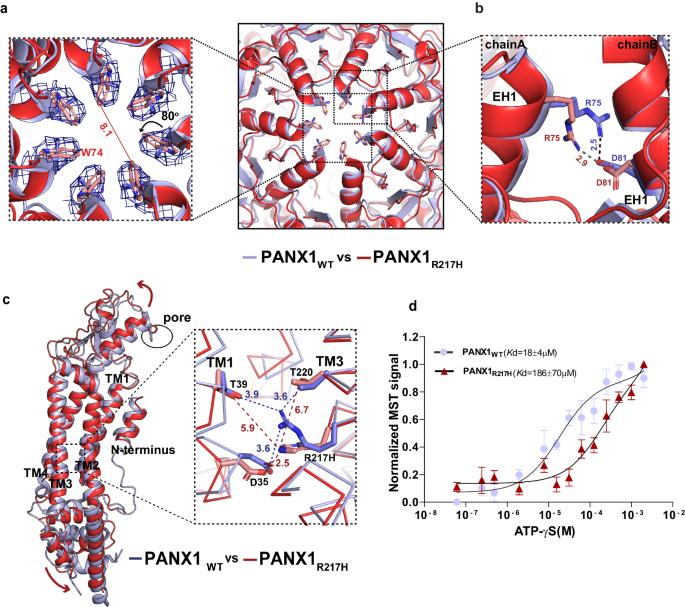
a A cross-section of superposed structures of PANX1WT (blue) and PANX1R217H (red); the inset shows the altered conformation of the residue (W74) at the extracellular entrance of the pore. The dashed line represents the reduced pore cross-section distance in PANX1R217H(red); the W74 rotates by 80° at the Ï2 torsion angle, reducing the pore diameter by 3.8âà compared to PANX1WT (Supplementary Movie 1). The density corresponding to W74, depicted at 5.0 Ï, is illustrated in the PANX1R217H mutant channel. b The residue R75 in PANX1R217H forms a salt bridge interaction with D81 of the adjacent subunit, mirroring the interaction observed in PANX1WT. c The structural superposition of PANX1WT and PANX1R217H exhibits a disrupted hydrogen-bond network, owing to the mutation, displayed in the inset. For clarity, only one subunit is shown, and arrows indicate the direction of the movement of the mutant in comparison to PANX1WT. The distances displayed in the figure are in angstroms (à ). d Weak apparent binding affinity of the PANX1R217H with ATP-γs was determined to be 186â±â70âµM compared to 18â±â4âµM of PANX1WT, nâ=â3 independent experiments, error bar represents S.D.
We investigated the residue environment in the vicinity of R217 within a 4âà radius in TM3 and observed hydrogen bond interaction with D35 in TM1 (3.6âà ) in PANX1WT. R217 also interacts with T220 in TM3 and T39 in TM1 through hydrogen bonds (Fig. 5c). Mutating arginine to histidine disrupts the H-bond interaction network within TMs 1 and 3. The H-bond interaction of T220 in TM3 and T39 in TM1 with R217H is disrupted due to the shorter side chain of histidine compared to arginine. The G44 in TM1 that likely acts as a hinge point facilitates the TM1 bending and displacement as a consequence of R217H substitution. The displacement observed in TM1 translates to an outward movement of ECD that could facilitate greater flexibility of the pore lining residue W74 to constrict the channel (Fig. 5a).
In comparison to the PANX1WT, the modeled side chain W74 has a Ï2 torsion angle shift of nearly 80° towards the pore leading to a constriction of the pore diameter in the mutant PANX1 (Fig. 5a, c). The current density for PANX1WT was twofold higher than PANX1R217H, at a positive voltage of 100âmV which could be associated with the partial closure of the pore (Fig. 4e, Supplementary Fig. 8a). Although there was a significant decrease in the current density in the R217H mutant, the mutation did not have significant effect on the CBX binding, as we could observe the inhibition of PANX1R217H currents by CBX similar to PANX1WT (Supplementary Fig. 7d). A comparison of the conductance density (nS/pF) to voltage reveals highly weakened channel conductance in response to increasing voltage in comparison to PANX1WT (Supplementary Fig. 8c). A comparison of normalized conductance to voltage for PANX1R217H and PANX1WT reveals altered voltage sensitivity (V50) of the mutant channel from around 40âmV for the PANX1WT to over 100âmV indicating reduced voltage sensitivity as a consequence of R217H mutation (Fig. 4f, g).
Binding studies using MST display a significant decrease in ATP-γS binding in the PANX1R217H mutant compared to PANX1WT suggesting that mutant behaves differently than the PANX1WT and have reduced ability to bind to ATP-γS (Fig. 5d). Although the R217H substitution is in the TM3, we detect structural shifts in the ECD indicating allosteric effects of this germline mutant. Such allosteric effects are observed in the case of disease-causing mutants where the mutation site is far from the observed structural changes and affects their functional properties39. It was proposed in an earlier study that the PANX1R217H interactions with the C-terminus can cause the altered channel properties in this germline mutant40. Given the allosteric effects of R217H on the pore diameter observed in this study, long-range effects in C-terminus could also drive some of the properties observed with PANX1R217H. However, given the absence of a structured C-terminus in PANX1 structures determined thus far, the influence of the C-terminus on PANX1R217H structure would be speculative at this juncture.