Participants
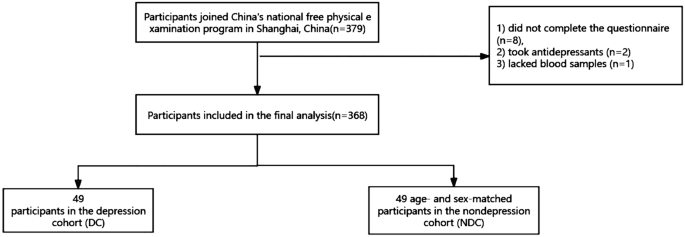
All of the subjects were individuals agedââ¥â65 who n. This study included 379 subjects who were invited to complete a comprehensive geriatric assessment and a face-to-face interview in the local community hospital. Our questionnaire assessed sociodemographic, lifestyle and health information. Sociodemographic variables included age and sex. Lifestyle includes smoking, drinking and daily activity levels. Daily activity levels were measured using the short form of the International Physical Activity Questionnaire (IPAQ)18. Health information included BMI, chronic conditions (such as diabetes, hypertension, hyperlipidemia, stroke, and heart disease, medication use and cognitive function. Cognitive function was assessed by the Mini-Mental State Examination (MMSE)19. Details of the questionnaire have been described in our previous study20. We excluded subjects who (1) did not complete the questionnaire (nâ=â8), (2) took antidepressants (nâ=â2) and (3) lacked blood samples (nâ=â1). Our subject screening process is shown in Fig. 1. The protocol of our study was reviewed and approved by the ethics committee at Shanghai University of Medicine and Health Sciences, China, and the methods were carried out in accordance with the principles of the Declaration of Helsinki. All the subjects provided informed consent before participation.
A flowchart of participant selection.
Measures of depression
Depression was measured using the 30-item geriatric depression scale (GDS)21. On this scale, items 2â4, 6, 8, 10â14, 16â18, 20, 22â26, and 28 are scored 1 point if answered âyesâ, and items 1, 5, 7, 9, 15, 19, 21, 27, 29 and 30 are scored 1 point if answered ânoâ. A total score of more than 10 points was considered to indicate depression. According to our standard, there were 49 subjects in the depression cohort (DC) and 49 age- and sex-matched individuals in the non-depression cohort (NDC).
Sample collection and preparation
Sample collection and LCâMS have been described in detail in our previous study19. Each plasma sample was collected from the study subjects on an empty stomach in the morning and stored at ââ80 °C until analysis. Before LCâMS, 150 μl of plasma that was thawed at room temperature was added to a 1.5-ml Eppendorf tube with 10 μl of 2-chlorophenylalanine (0.3 mg/ml) dissolved in methanol as an internal standard and 450 μl of a mixture of methanol/acetonitrile (2/1) to remove the protein and then vortexed for 1 min. The mixture was extracted by ultrasonication for 10 min, stored for 30 min (ââ20â) and then centrifuged at 4 °C for 10 min (13,000 rpm). Two hundred microlitres of supernatant was dried in a freeze concentration centrifuge dryer, redissolved in 300 μl of methanol/water (1/4), vortexed for 30 s, and extracted by ultrasonication for 3 min. The sample was centrifuged at 4 °C for 10 min (13,000 rpm), and 150 μl of supernatant was filtered through 0.22-μm microfilters and transferred to LC vials. The vials were stored atâââ80 °C until LCâMS.
The pretreatment for GCâMS was similar to that for LCâMS. A total of 150 μl of plasma was added to an Eppendorf tube with 20 μl of 2-chlorophenylalanine (0.3 mg/ml) dissolved in methanol as an internal standard and vortexed for 10 s. Then, 450 μl of an ice-cold mixture of methanol/acetonitrile (2/1, v/v) to remove the protein was added to the tube and vortexed for 30 s. The mixture was extracted by ultrasonication in an ice water bath for 10 min, stored for 30 min (ââ20 °C), and centrifuged at 4 °C for 10 min (13,000 rpm). Two hundred millilitres of supernatant was placed into a new glass bottle, dried in a freeze concentration centrifuge and added to 80 μL of 15 mg/mL methoxylamine hydrochloride in pyridine. The resultant mixture was vortexed for 2 min and incubated at 37 °C for 90 min. Then, 50 μL of BSTFA (with 1% TMCS) and 20 μL of n-hexane were added into the bottle, and the bottle was vortexed violently for 2 min and derivatized at 70 °C for 60 min. The samples were placed at room temperature for 30 min before GCâMS.
LCâMS and GCâMS
LCâMS was performed on the ACQUITY UPLC I-Class system (Waters Corporation, Miford, USA) coupled with VION IMS QT of the high-resolution mass spectrometer (Waters Corporation, Milford, USA). An ACQUITY UPLC BEH C18 column (1.7 μm, 2.1âÃâ100 mm) was employed in both the positive and negative models. GCâMS was performed on an Agilent 7890B gas chromatography system coupled to an Agilent 5977A MSD system (Agilent Technologies Inc., CA, USA). A DB-5MSf used-silica capillary column (30 mâÃâ0.25 mmâÃâ0.25 μm, Agilent J& W Scientific, Folsom, CA, USA) was utilized to separate the derivatives. To monitor the stability and repeatability of LCâMS and GCâMS, QC samples were inserted regularly and analysed in every ten samples.
Metabolite identification and analysis
The LCâMS data were analysed using Proggenesis Qi software version 2.3 (Nonlinear, Dynamics, Newcastle, UK). First, the software is used to carry out meaningful data mining and perform advanced alignment, picking, normalization, and retention time (RT) correction. The obtained characteristic matrix includes information about the mass charge ratio (m/z), RT, and peak intensities. Then, the identification of metabolites was based on precise m/z, secondary fragments, and isotope distribution using the human metabolome database (HMDB), Human Metabolome Database (HMDB) (http://www.hmdb.ca/), lipid maps (version 2.3) (http://www.lipidmaps.org/), METLIN (http://metlin.scripps.edu/), and self-built databases (EMDB) for qualitative analysis.
The GCâMS data used the software MS-DIAL version 2.74 for peak detection, peak identification, characterization, peak alignment, wave filtering, etc. Metabolites were annotated through the LUG database (Untargeted database of GCâMS rom Lumingbio). The raw data matrix was obtained from the raw data with a three-dimensional dataset, including sample information, the name of the peak of each substance, retention time, retention index, mass-to-charge ratio, and signal intensity, after alignment with the Statistical Compare component. The internal standards with RSDâ>â0.3 were used to segment and normalize all peak signal intensities in each sample, and the segmented and normalized results were removed redundancy and merged peak to obtain the data matrix.
A total of 1008 compound identifications detected by LCâMS and 446 compound identifications detected by GCâMS were automatically linked to the compounds. Finally, orthogonal partial least-squares discriminant analysis (OPLS-DA) was used to visualize the differences in metabolites between DC and NDC, and 200 response permutation tests (RPTs), including parameters such as R2 and Q2, were used to quantify the goodness of fit and assess the reliability of the established models. If these parameters were close to 1.0, the model was considered valid. Multidimensional coupling and single-dimensional analysis were used to select different metabolites between groups. The variable importance in projection (VIP) generated in OPLS-DA represented differential metabolites with biological significance. Furthermore, the significance of differential metabolites was further verified by Studentâs t test. Variables with VIPâ>â1.0 and pâ<â0.05 were considered to be differential metabolites. To quantify the diagnostic performance of differential metabolites, a receiver operating characteristic curve (ROC) analysis was carried out, and the value of the area under the ROC curve (AUC) was calculated.
Pathway analysis
To determine the mechanism of metabolic pathway variation, the differential metabolites were based on the Kyoto Encyclopedia of Genes and Genomes (KEGG) database (http://www.kegg.jp/kegg/pathway.html) to carry out metabolic pathway enrichment analysis. Their KEGG ID and pathway were found, and then the number of metabolites enriched in the corresponding pathway was calculated. The pathway with a pâ<â0.05 was selected as an enriched pathway; its calculation formula is given as follows:
$$P = \mathop \sum \limits_{i = 0}^{m – 1} \frac{{\left( \frac{M}{i} \right)\left( {\frac{N – M}{{n – i}}} \right)}}{\frac{N}{n}}$$
where N is the total number of metabolites, n is the number of differential metabolites, M is the number of metabolites annotated as a specific pathway, and m is the number of differential metabolites annotated as a specific pathway.
Statistical analyses
Baseline sociodemographic and health-related characteristic analyses were performed using SPSS version 25.0 (SPSS Incorporation, Chicago, IL, USA), and pâ<â0.05 was regarded as statistically significant. Baseline sociodemographic and health-related characteristics were compared between the DC and the NDC using an independent t test for numeric variables and a chi-square test for categorical variables. Data with a normal distribution are expressed as the meanâ±âSD, and categorical variables are expressed as proportions.